Importance of Binding Site Hydration and Flexibility Revealed When Optimizing a Macrocyclic Inhibitor of the Keap1-Nrf2 Protein-Protein Interaction.
Begnini, F., Geschwindner, S., Johansson, P., Wissler, L., Lewis, R.J., Danelius, E., Luttens, A., Matricon, P., Carlsson, J., Lenders, S., Konig, B., Friedel, A., Sjo, P., Schiesser, S., Kihlberg, J.(2022) J Med Chem 65: 3473-3517
- PubMed: 35108001
- DOI: https://doi.org/10.1021/acs.jmedchem.1c01975
- Primary Citation of Related Structures:
7Q5H, 7Q6Q, 7Q6S, 7Q8R, 7Q96 - PubMed Abstract:
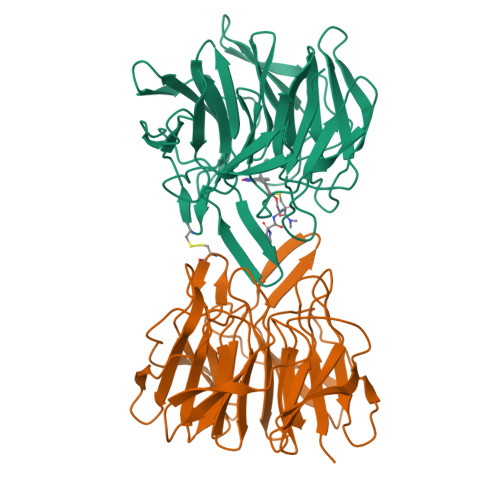
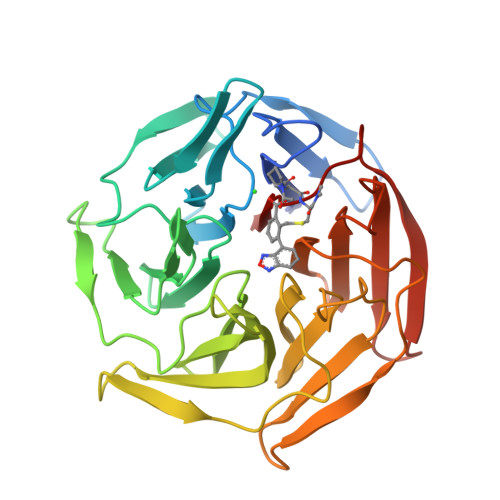
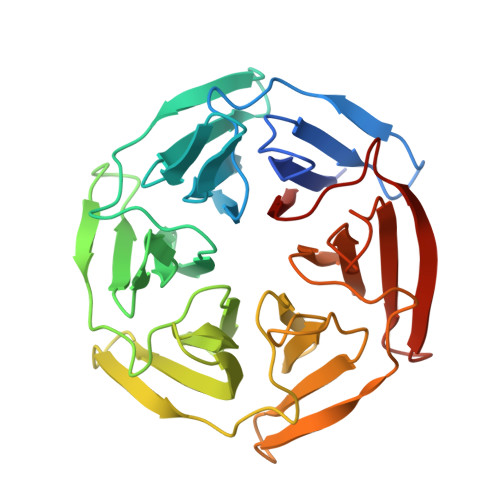
Upregulation of the transcription factor Nrf2 by inhibition of the interaction with its negative regulator Keap1 constitutes an opportunity for the treatment of disease caused by oxidative stress. We report a structurally unique series of nanomolar Keap1 inhibitors obtained from a natural product-derived macrocyclic lead. Initial exploration of the structure-activity relationship of the lead, followed by structure-guided optimization, resulted in a 100-fold improvement in inhibitory potency. The macrocyclic core of the nanomolar inhibitors positions three pharmacophore units for productive interactions with key residues of Keap1, including R415, R483, and Y572. Ligand optimization resulted in the displacement of a coordinated water molecule from the Keap1 binding site and a significantly altered thermodynamic profile. In addition, minor reorganizations of R415 and R483 were accompanied by major differences in affinity between ligands. This study therefore indicates the importance of accounting both for the hydration and flexibility of the Keap1 binding site when designing high-affinity ligands.
Organizational Affiliation:
Department of Chemistry─BMC, Uppsala University, Box 576, 75123 Uppsala, Sweden.